Building a human tumour microenvironment in the lab – updates from the CanBuild Project

Researchers from Barts Cancer Institute at Queen Mary University of London, led by Professor Fran Balkwill and Dr Oliver Pearce, have built two 3D multi-cellular models of the human tumour microenvironment (TME) in ovarian cancer. The models, which are the first created from the CanBuild project, have revealed novel insights into the role of the TME in cancer progression.
Tumours are not just a mass of cancerous cells; they also contain healthy cells that have been corrupted by the cancer to help it grow and spread. The cancer cells and corrupted healthy cells, along with structural molecules that support the tumour (the extracellular matrix or ECM), constitute what is known as the TME.
Back in 2013, Professor Balkwill assembled a team of cancer researchers, bioengineers and tissue engineers for the CanBuild project. Funded by the European Research Council (ERC), the aim of the project was to build mini models of the human TME in the laboratory to study more accurately how the TME contributes to cancer development and progression, and to provide a platform to test new treatments.
Focussing on a common site of cancer spread (metastasis) in ovarian cancer called the omentum, the team first deconstructed human omental tumour samples to characterise the different cell types (and their proportions) present in the TME. This deconstruction provided the building blocks needed to re-construct 3D multi-cellular cancer models in the laboratory, using cells obtained from cancer biopsies.
Two recent publications published in iScience describe two models created by the team. By adding cell types into each model one by one, the team have been able to understand the roles of specific cell types within the TME, and learn more about ovarian cancer cell invasion and the regulation of molecules involved in cancer progression.
Dr Pearce said:
“Building on our deconstruction work, we have been able to build 3D multi-cellular models that can recapitulate the complex interactions between cancer cells and their microenvironment. These models have allowed us to answer important scientific questions about the role of the TME in ovarian cancer. Investigations such as these would not be possible in 2D models or animal models.”
Professor Balkwill added:
“We are very excited about the potential of our models and believe they are a useful first step in pre-clinical evaluation of therapies targeting the TME in human solid cancers.”
The published research was funded by the ERC and Cancer Research UK, and performed in collaboration with researchers from Queen Mary’s School of Engineering and Materials Science.
Platelets are a key player in cancer cell invasion
Cancer cell invasion is a key process in progression of the disease, as it allows the cancer cells to escape the primary tumour and enter the bloodstream, where they may travel to and establish tumours in other sites in the body.
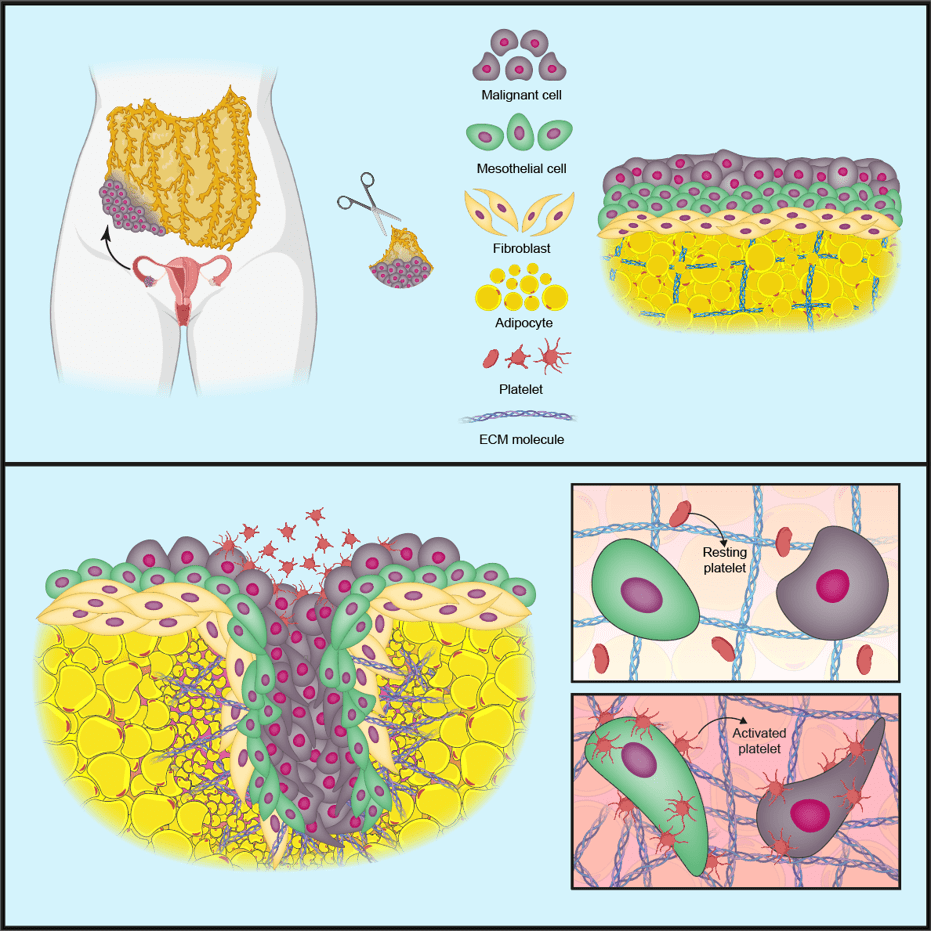
In the first iScience publication, prompted by previous research and findings in patient biopsies, the team created a model to investigate the role of blood cells called platelets in cancer cell invasion and production of the tumour-supporting ECM.
The team first made a model of a normal omentum using three cell types taken from human omental tissue samples: mesothelial cells (form a protective barrier on tissues), fibroblasts (cells involved in making the ECM) and fat cells called adipocytes. They then added in cancer cells from cell lines of the most common type of ovarian cancer, high-grade serous ovarian cancer (HGSOC).
In this ‘tetra-culture’ model, the cancer cells attached themselves to mesothelial cells on the surface of the tissue; however, when the team added human platelets to the model the cancer cells began to invade into the surrounding tissue.
Further investigations revealed that the platelets attach to the mesothelial cell barrier on the surface of the model, switching the platelets to an ‘activated’ state. The activated platelets change the shape and behaviour of the mesothelial cells, causing the mesothelial barrier to break down and allowing the cancer cells to invade into the underlying tissue.
The activated platelets also increased the production of proteins that change or remodel the surrounding ECM. ECM remodelling is associated with poor prognosis in HGSOC, as it creates an environment that supports cancer spread.
Modelling features linked to poor cancer prognosis
The team’s previous deconstruction work identified a pattern of 22 genes, termed the matrix index (MI), which changed overtime as cancer progressed. The MI could distinguish patients with shorter overall survival in HGSOC and 12 other solid tumour types. Six MI genes in particular were found to be significantly upregulated in cancer, causing the increased production of ECM proteins associated with tumour progression, poor prognosis, and cancer cell invasion.
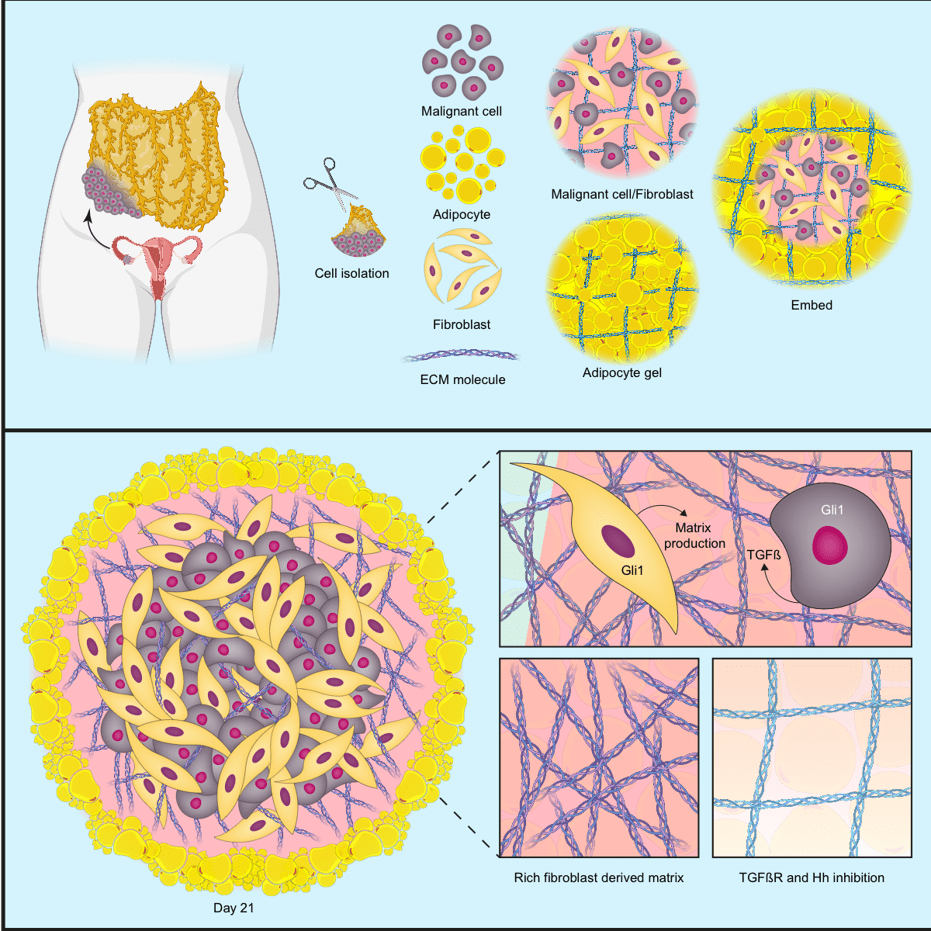
In the second iScience publication, the team set out to make a 3D human multi-cellular model of HGSOC metastasis that could capture the cells and signalling pathways in the TME that control the production of the six upregulated MI proteins.
Starting with analyses in patient biopsies, the team found that the upregulated proteins were released from both cancer cells and normal cells within the TME, and that molecular messengers called TGFβ and the Hedgehog pathway were important regulators of these proteins. To investigate these associations further, the team constructed a model made up of three cell types: adipocytes and fibroblasts taken from human omental tissue samples, and cancer cells from HGSOC cell lines.
The model successfully recapitulated the key features associated with poor prognosis in HGSOC; when the three cell types were put together, they generated all six MI proteins. The model showed that the cancer cells released TGFβ, which triggered the fibroblasts to remodel the ECM via Hedgehog signalling, creating an environment that supported the production of the MI proteins.
A platform for testing potential treatments
In both models, TGFβ was a major player contributing to cancer progression, suggesting that it may be an effective target for cancer therapies. By adding an inhibitor that blocked the effects of TGFβ into the platelet model, the team were able to disrupt platelet activation and stop invasion of cancer cells into the tissue. In the second model, the addition of a combination of TGFβ and Hedgehog inhibitors stopped the fibroblasts from remodelling the ECM and prevented the production of the MI molecules associated with poor prognosis.
The models have allowed the team to dissect the roles of different cell types within the TME of human HGSOC, and identify the processes governing disease progression in ways that would not be possible in animal models. The findings suggest that blocking interactions between key cell types in the TME via messenger molecules, such as TGFβ, may have clinical potential, and that the 3D multi-cellular models represent a useful pre-clinical platform for testing therapies targeted at the TME. Other cells, such as immune cells called macrophages, can also be added to these models, increasing their ability to test novel biological therapies.
Category: General News, Publications

No comments yet